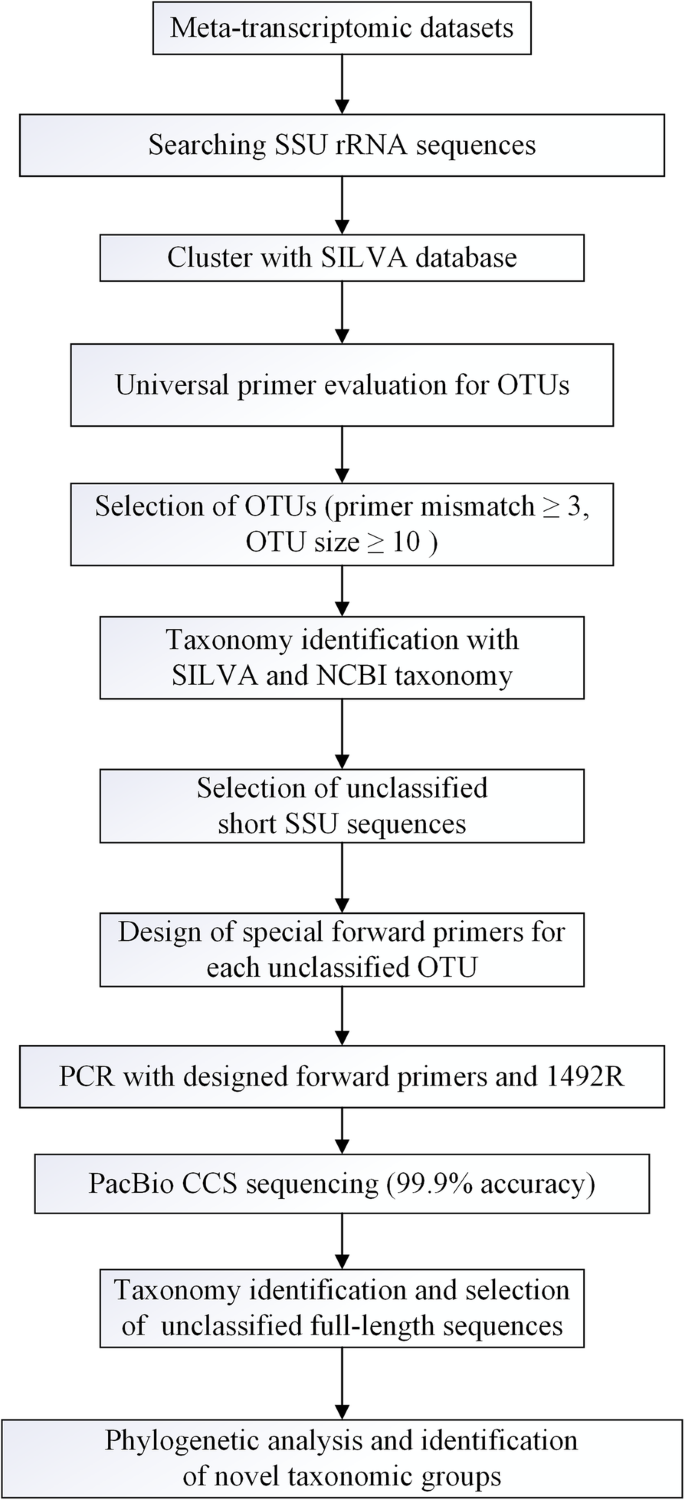
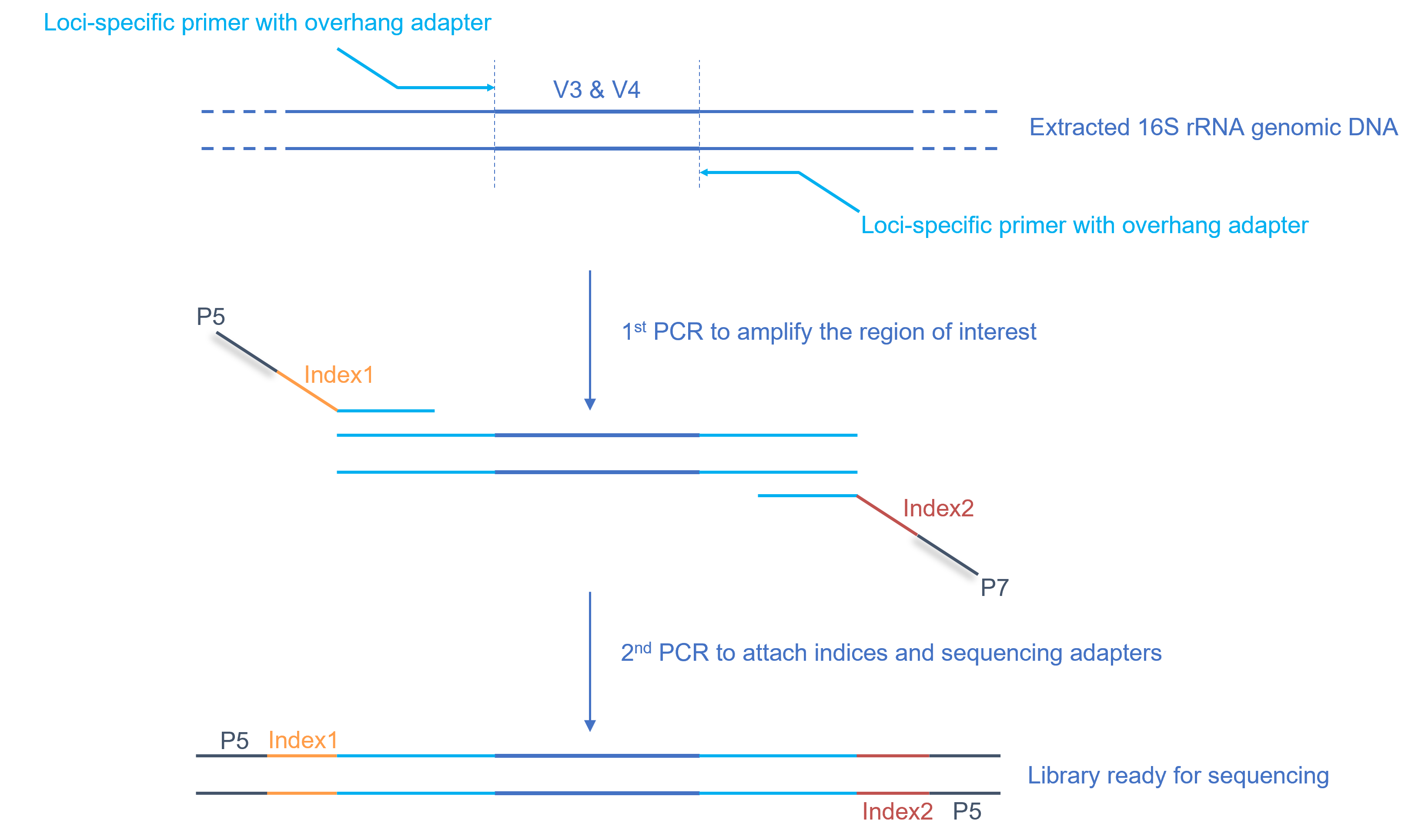
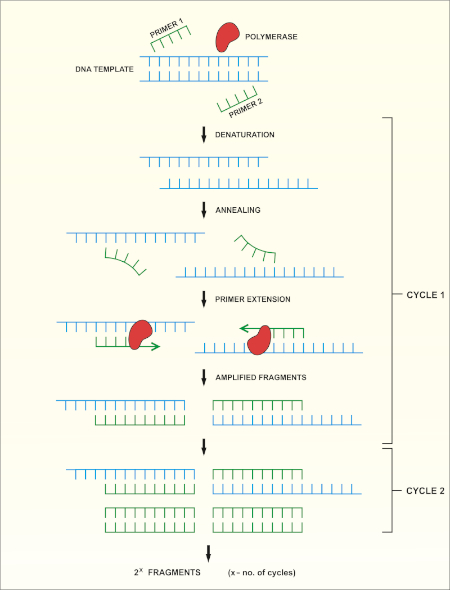
Absolute quantitation of microbes using 16S rRNA gene metabarcoding: A rapid normalization of relative abundances by quantitative PCR targeting a 16S rRNA gene spike‐in standard - Zemb - 2020 - MicrobiologyOpen - Wiley Online Library
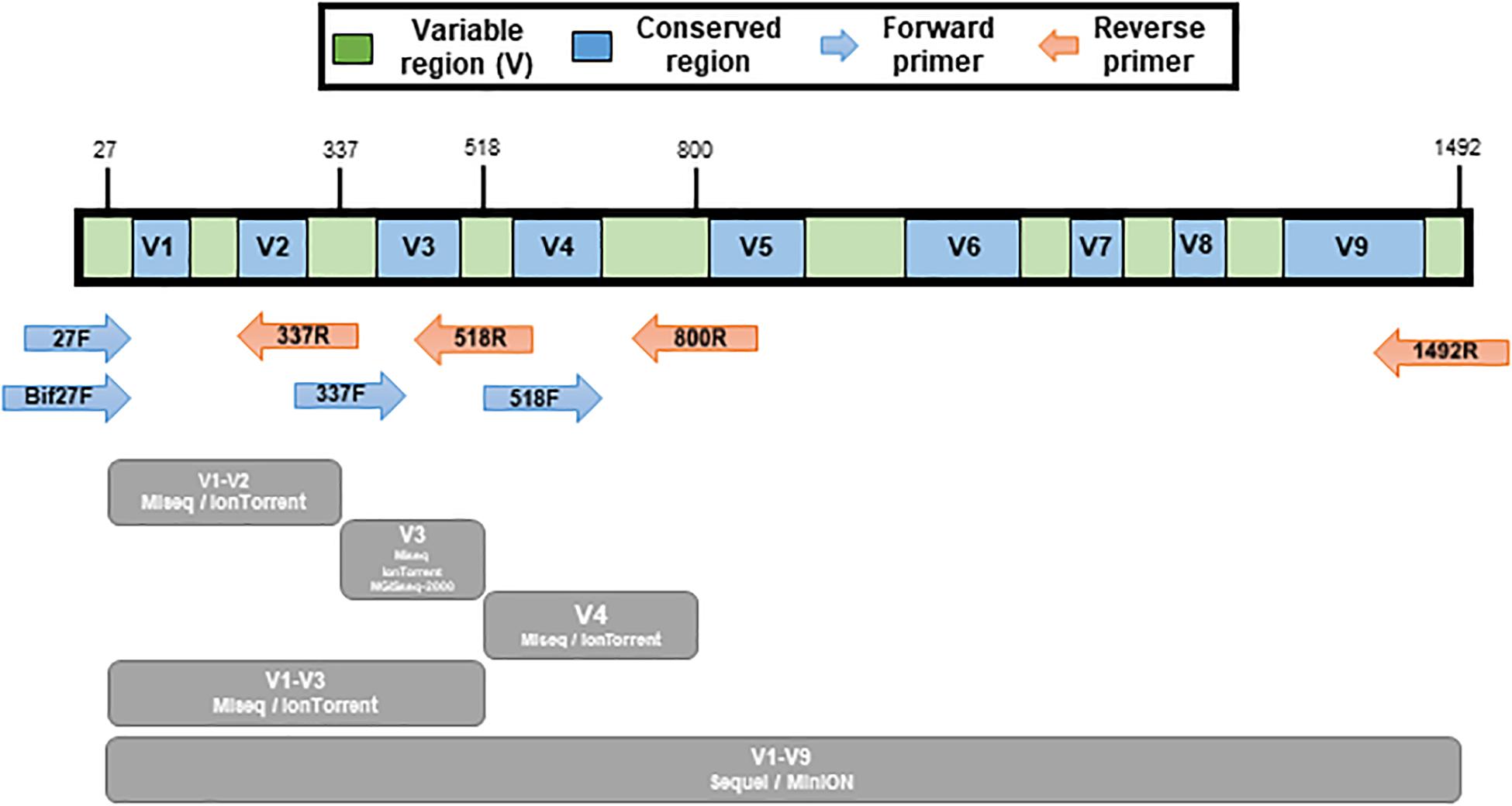
Frontiers | Comparison of 16S rRNA Gene Based Microbial Profiling Using Five Next-Generation Sequencers and Various Primers
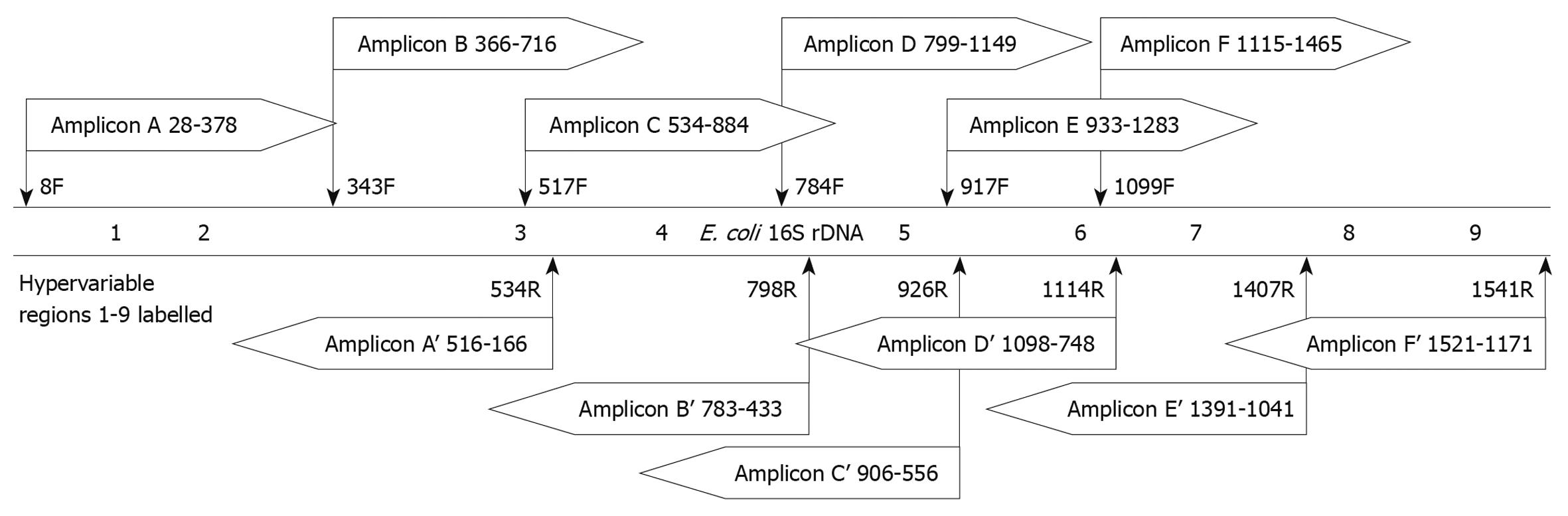
![PDF] Conservative Fragments in Bacterial 16S rRNA Genes and Primer Design for 16S Ribosomal DNA Amplicons in Metagenomic Studies | Semantic Scholar PDF] Conservative Fragments in Bacterial 16S rRNA Genes and Primer Design for 16S Ribosomal DNA Amplicons in Metagenomic Studies | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/efdba25b13c217b52790c95a5a7d23b4537fc70e/2-Table1-1.png)
PDF] Conservative Fragments in Bacterial 16S rRNA Genes and Primer Design for 16S Ribosomal DNA Amplicons in Metagenomic Studies | Semantic Scholar
Biased Diversity Metrics Revealed by Bacterial 16S Pyrotags Derived from Different Primer Sets | PLOS ONE

Selecting 16S rRNA primers for microbiome analysis in a host-microbe system: the case of the jellyfish Rhopilema nomadica | bioRxiv
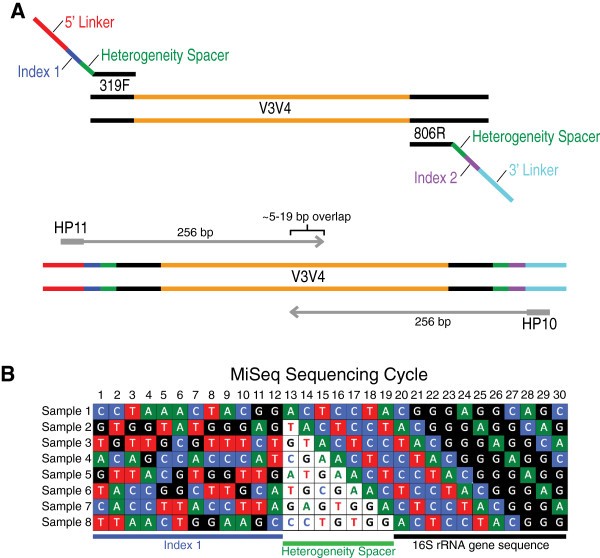
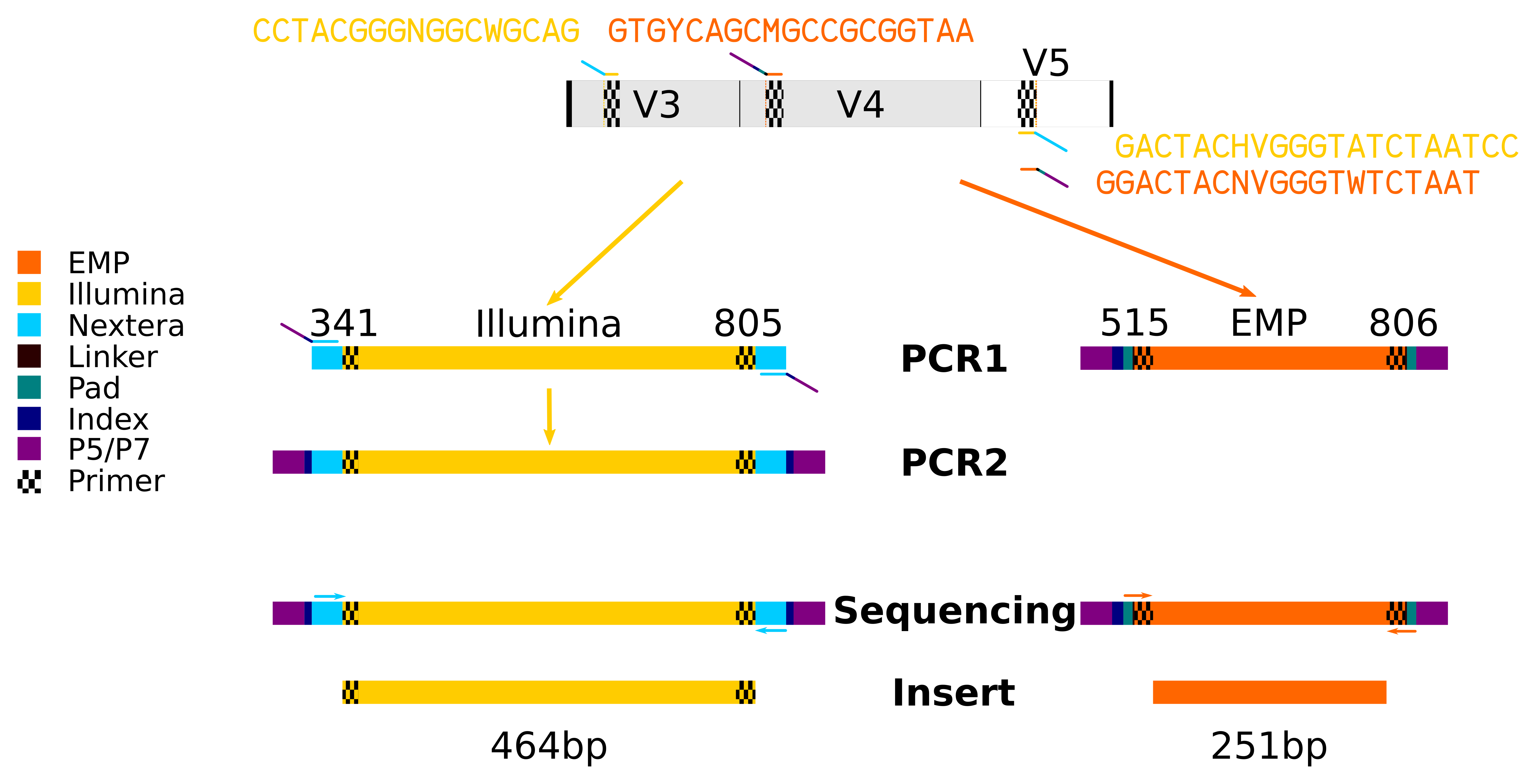
An improved dual-indexing approach for multiplexed 16S rRNA gene sequencing on the Illumina MiSeq platform | Microbiome | Full Text

Sequences of the forward (F) and reverse (R) primers, for 16S rRNA gene | Download Scientific Diagram
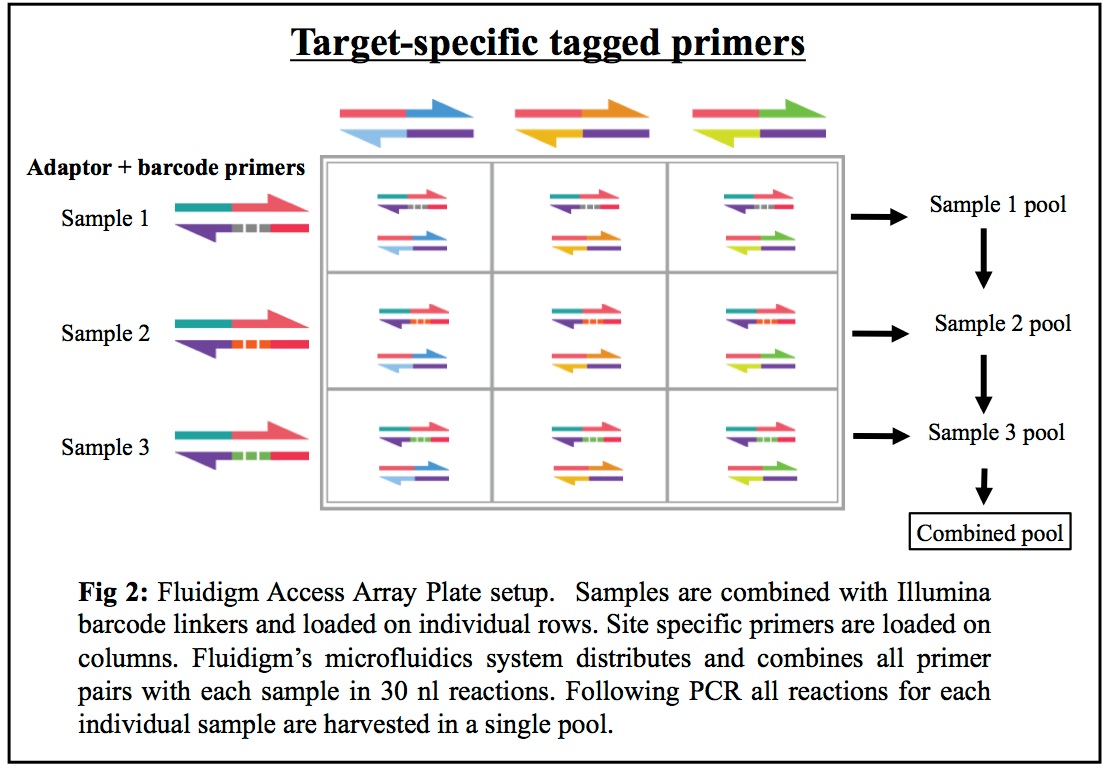
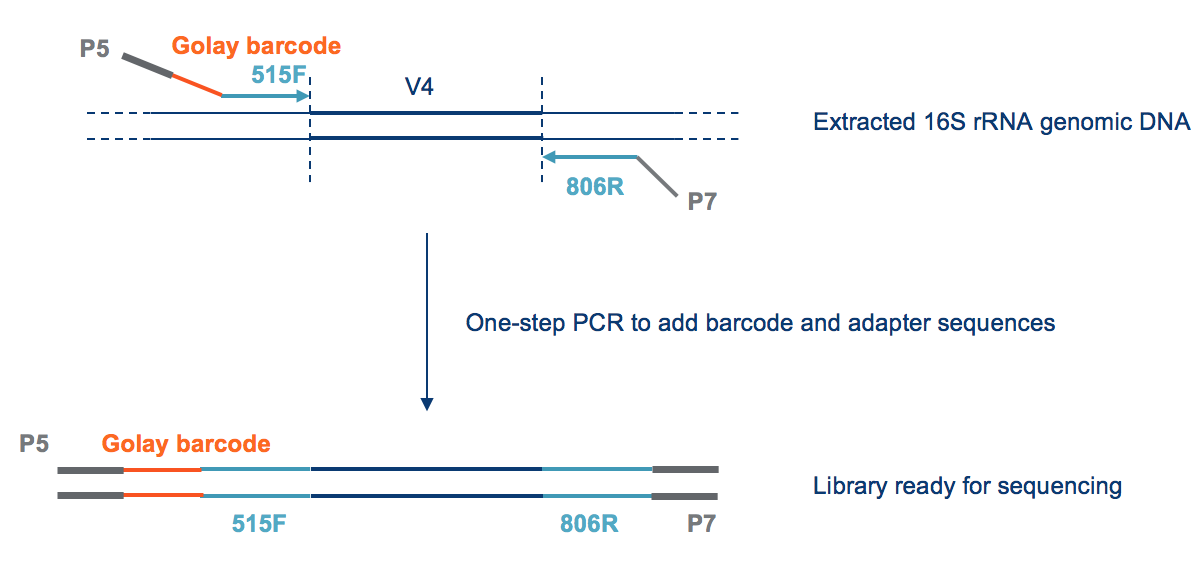
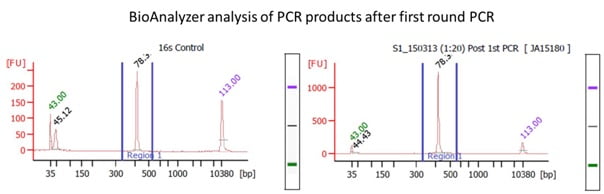
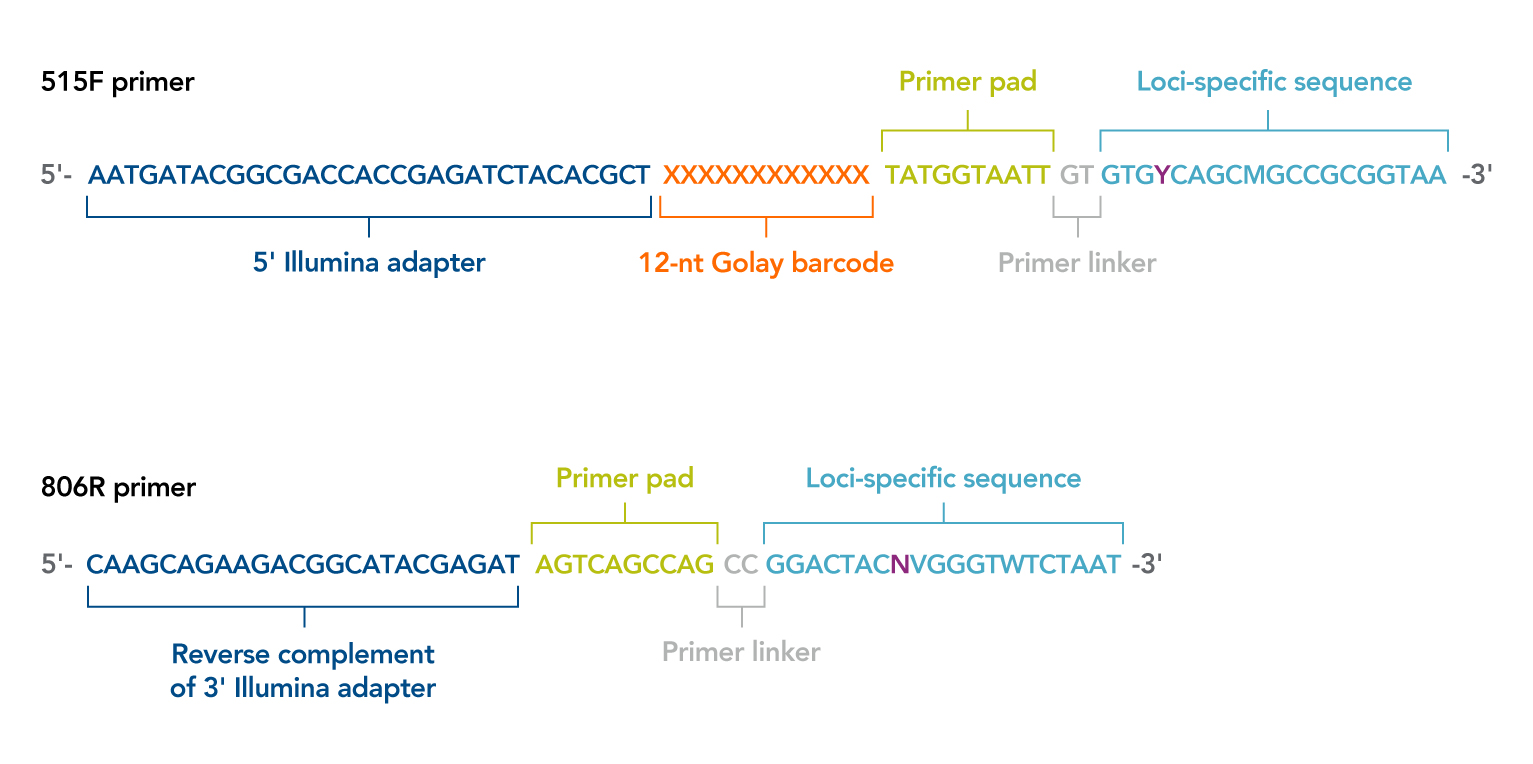
HTDNA: Services/Equipment: Preparation of 16S rRNA (and all other) Amplicons for Illumina Sequencing | Biotech

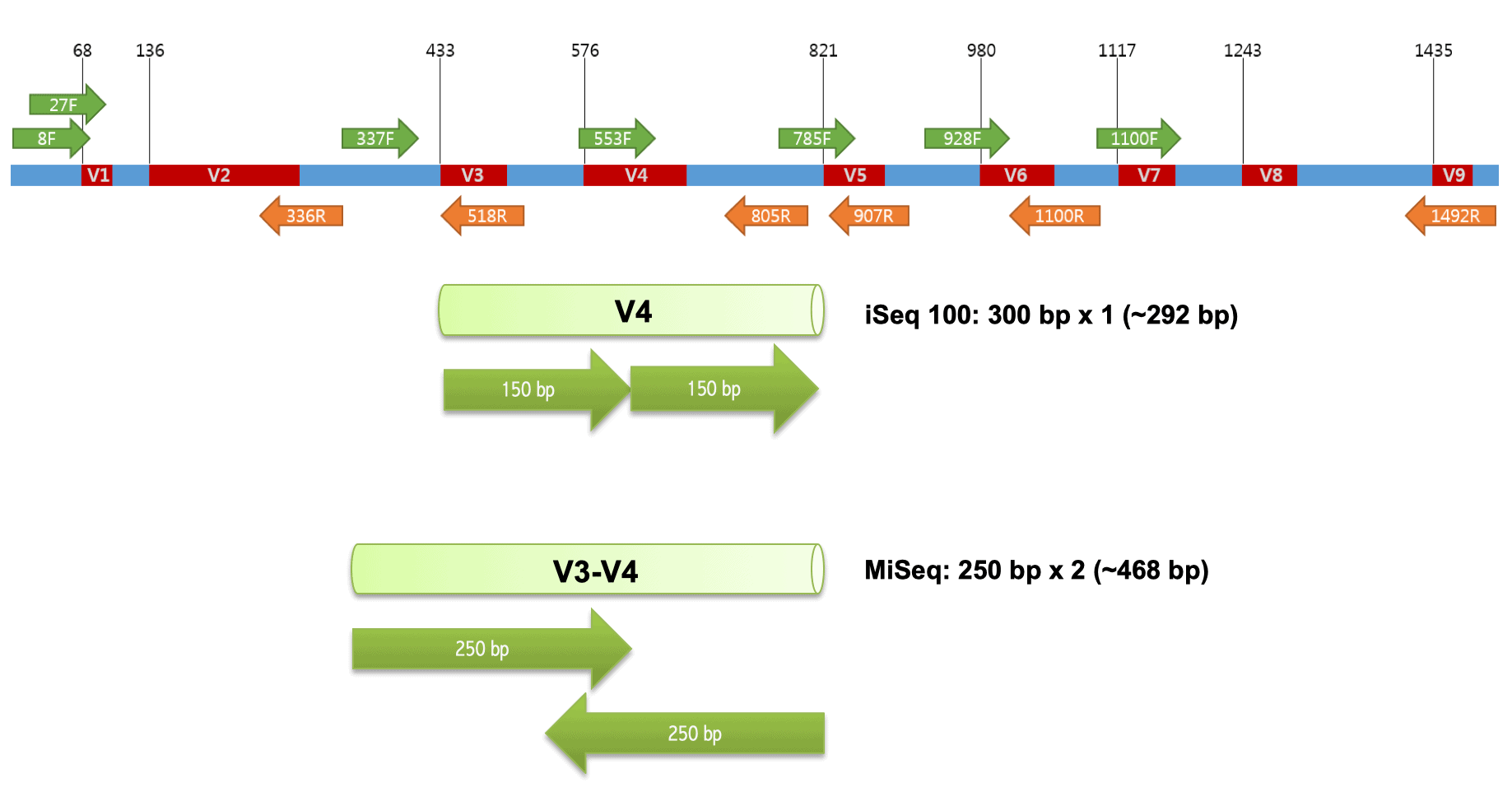
Better skin microbiome analyses using new 16S V4 region primers developed by Microbiome Insights' scientific team — Microbiome Insights